Introduction
The function bgmCompare() extends bgm() to
independent-sample designs. It estimates whether edge weights and
category thresholds differ across groups in an ordinal Markov random
field (MRF).
Posterior inclusion probabilities indicate how plausible it is that a group difference exists in a given parameter. These can be converted to Bayes factors for hypothesis testing.
Fitting a model
fit = bgmCompare(x = data_adhd, y = data_no_adhd, seed = 1234)Posterior summaries
The summary shows both baseline effects and group differences:
summary(fit)
#> Posterior summaries from Bayesian grouped MRF estimation (bgmCompare):
#>
#> Category thresholds:
#> parameter mean mcse sd n_eff Rhat
#> 1 avoid (1) -2.570 0.012 0.389 1000.548 1.001
#> 2 closeatt (1) -2.255 0.011 0.372 1058.581 1.013
#> 3 distract (1) -0.450 0.013 0.330 650.281 1.000
#> 4 forget (1) -1.540 0.011 0.326 860.980 1.002
#> 5 instruct (1) -2.334 0.014 0.394 790.011 1.005
#>
#> Pairwise interactions:
#> parameter mean mcse sd n_eff Rhat
#> 1 avoid-closeatt 0.811 0.017 0.445 700.771 1.002
#> 2 avoid-distract 1.622 0.012 0.372 1010.162 1.000
#> 3 avoid-forget 0.478 0.012 0.363 846.177 1.000
#> 4 avoid-instruct 0.381 0.014 0.431 989.191 1.003
#> 5 closeatt-distract -0.168 0.011 0.359 1155.071 1.006
#> 6 closeatt-forget 0.153 0.008 0.293 1373.018 1.001
#> ... (use `summary(fit)$pairwise` to see full output)
#>
#> Inclusion probabilities:
#> parameter mean mcse sd n0->0 n0->1 n1->0 n1->1
#> avoid (main) 1.000 0.000 0 0 0 1999
#> avoid-closeatt (pairwise) 0.696 0.018 0.460 397 210 211 1181
#> avoid-distract (pairwise) 0.386 0.012 0.487 803 425 424 347
#> avoid-forget (pairwise) 0.861 0.014 0.346 165 113 113 1608
#> avoid-instruct (pairwise) 0.996 0.002 0.067 4 5 5 1985
#> closeatt (main) 1.000 0.000 0 0 0 1999
#> n_eff_mixt Rhat
#>
#> 662.308 1.001
#> 1623.275 1
#> 618.031 1.001
#> 774.056 1.126
#>
#> ... (use `summary(fit)$indicator` to see full output)
#> Note: NA values are suppressed in the print table. They occur when an indicator
#> was constant or had fewer than 5 transitions, so n_eff_mixt is unreliable;
#> `summary(fit)$indicator` still contains all computed values.
#>
#> Group differences (main effects):
#> parameter mean mcse sd n_eff n_eff_mixt Rhat
#> avoid (diff1; 1) -2.585 0.738 822.680 1.001
#> closeatt (diff1; 1) -2.884 0.719 850.621 1.000
#> distract (diff1; 1) -2.592 0.671 588.723 1.000
#> forget (diff1; 1) -2.878 0.642 646.747 1.002
#> instruct (diff1; 1) -2.386 0.893 521.021 1.000
#> Note: NA values are suppressed in the print table. They occur here when an
#> indicator was zero across all iterations, so mcse/n_eff/n_eff_mixt/Rhat are undefined;
#> `summary(fit)$main_diff` still contains the NA values.
#>
#> Group differences (pairwise effects):
#> parameter mean mcse sd n_eff n_eff_mixt
#> avoid-closeatt (diff1) 0.948 0.032 0.865 568.814 724.944
#> avoid-distract (diff1) 0.211 0.012 0.355 932.378 938.052
#> avoid-forget (diff1) 1.281 0.029 0.796 579.667 764.262
#> avoid-instruct (diff1) -2.745 0.034 0.990 782.659 827.328
#> closeatt-distract (diff1) -0.159 0.012 0.324 974.294 741.833
#> closeatt-forget (diff1) 0.164 0.012 0.324 1029.198 755.929
#> Rhat
#> 1.004
#> 1.001
#> 1.000
#> 1.003
#> 1.007
#> 1.001
#> ... (use `summary(fit)$pairwise_diff` to see full output)
#> Note: NA values are suppressed in the print table. They occur here when an
#> indicator was zero across all iterations, so mcse/n_eff/n_eff_mixt/Rhat are undefined;
#> `summary(fit)$pairwise_diff` still contains the NA values.
#>
#> Use `summary(fit)$<component>` to access full results.
#> See the `easybgm` package for other summary and plotting tools.You can extract posterior means and inclusion probabilities:
coef(fit)
#> $main_effects_raw
#> baseline diff1
#> avoid(c1) -2.5698070 -2.585211
#> closeatt(c1) -2.2549973 -2.884102
#> distract(c1) -0.4500195 -2.592396
#> forget(c1) -1.5397695 -2.878431
#> instruct(c1) -2.3342186 -2.385729
#>
#> $pairwise_effects_raw
#> baseline diff1
#> avoid-closeatt 0.8106540 0.9480407
#> avoid-distract 1.6217536 0.2109026
#> avoid-forget 0.4775235 1.2813476
#> avoid-instruct 0.3808413 -2.7450407
#> closeatt-distract -0.1679917 -0.1589278
#> closeatt-forget 0.1531440 0.1642686
#> closeatt-instruct 1.4875764 0.4870774
#> distract-forget 0.3755528 0.2333754
#> distract-instruct 1.1930973 1.2431688
#> forget-instruct 1.0594628 0.7473659
#>
#> $main_effects_groups
#> group1 group2
#> avoid(c1) -1.2772013 -3.862413
#> closeatt(c1) -0.8129461 -3.697048
#> distract(c1) 0.8461783 -1.746217
#> forget(c1) -0.1005540 -2.978985
#> instruct(c1) -1.1413540 -3.527083
#>
#> $pairwise_effects_groups
#> group1 group2
#> avoid-closeatt 0.33663370 1.2846744
#> avoid-distract 1.51630235 1.7272049
#> avoid-forget -0.16315030 1.1181973
#> avoid-instruct 1.75336163 -0.9916790
#> closeatt-distract -0.08852775 -0.2474556
#> closeatt-forget 0.07100968 0.2352783
#> closeatt-instruct 1.24403770 1.7311151
#> distract-forget 0.25886514 0.4922405
#> distract-instruct 0.57151284 1.8146817
#> forget-instruct 0.68577985 1.4331458
#>
#> $indicators
#> avoid closeatt distract forget instruct
#> avoid 1.0000 0.6960 0.3860 0.861 0.9955
#> closeatt 0.6960 1.0000 0.3835 0.392 0.5360
#> distract 0.3860 0.3835 1.0000 0.401 0.8335
#> forget 0.8610 0.3920 0.4010 1.000 0.7110
#> instruct 0.9955 0.5360 0.8335 0.711 1.0000Visualizing group networks
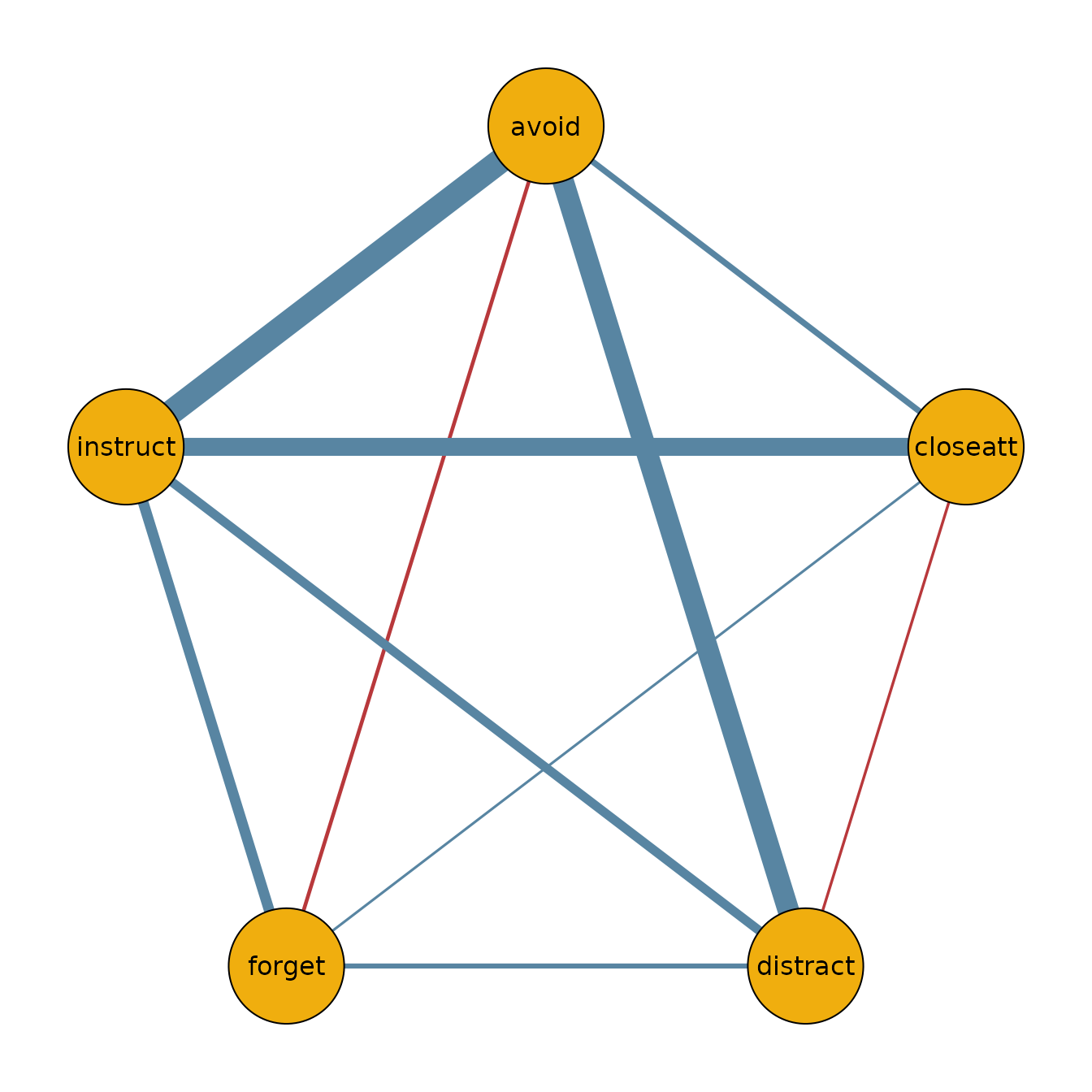
We can use the output to plot the network for the ADHD group:
library(qgraph)
adhd_network = matrix(0, 5, 5)
adhd_network[lower.tri(adhd_network)] = coef(fit)$pairwise_effects_groups[, 1]
adhd_network = adhd_network + t(adhd_network)
colnames(adhd_network) = colnames(data_adhd)
rownames(adhd_network) = colnames(data_adhd)
qgraph(adhd_network,
theme = "TeamFortress",
maximum = 1,
fade = FALSE,
color = c("#f0ae0e"), vsize = 10, repulsion = .9,
label.cex = 1, label.scale = "FALSE",
labels = colnames(data_adhd)
)